Work in the lab aims to integrate data from genomes and experiments to determine how complex host interactions shape bacterial evolution and ecology, and how these interactions feedback to impact hosts. We use a variety of systems to address these questions. Highlighted below are several ongoing projects from the lab.
Interactions of insects, plants and associated bacteria
We use a tractable system of interactions between the plant pathogen Pseudomonas syringae and plant and insect hosts. In addition to being a common plant pathogen, P. syringae is also a ubiquitous epiphyte on plant surfaces. It has recently been shown to grow to high numbers within some hemipteran insects, causing insect death. Hemipteran insects such as aphids are serious agricultural pests and are difficult to control with traditional methods like pesticides. We have found that some bacterial strains can cause infection in pea aphids with just a few cells and kill all infected aphids in as few as three days. These epiphytic bacteria may therefore have substantial impacts on insect pest populations, but the ecological and evolutionary implications of the interactions are largely unexplored.
Genetic basis for bacterial phenotypes that influence insects
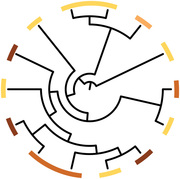
P. syringae is a phenotypically and genetically diverse group of strains with excellent genomic resources available. In work funded by the USDA NIFA, we are using genomic enabled methods such as RNA Seq and TnSeq to understand the genetic basis underlying the bacterial phenotypes of high virulence to insects, high infectivity, and ability to grow within insects. This study will analyze a breadth of strains within a phylogenetic context to understand how different strains kill insects and how this trait has evolved in bacterial populations. Postdoc Melanie Smee is currently leading work on this project.
Effects on insect hosts

Aphid response to infection
The lab is also interested to understand how aphids respond to and avoid highly virulent bacteria in the phyllosphere. Previous work has shown that insects may rely on non-immunological defenses such as symbiont mediated defense (found in sweet potato whiteflies infected with P. syringae) or fecundity compensation (found in pea aphids infected with P. syringae). A current project in the lab is focused on aphid avoidance of virulent bacteria. We have found that pea aphids can avoid some virulent strains in the phyllosphere, and therefore avoid infection. The goal of this project is to understand how aphids identify virulent bacteria and which bacterial traits are important.
Breadth of effects
In work funded by the USDA, we aim to understand how diverse aphid species are influenced by P. syringae, specifically how many aphid species can become infected by the bacteria, and how variables such as bacteria strain, aphid species, and plant host influence these interactions. We are focusing on aphid species that are local crop pests in hopes of understanding how these interactions may be used to control pests.
The lab is also interested to understand how aphids respond to and avoid highly virulent bacteria in the phyllosphere. Previous work has shown that insects may rely on non-immunological defenses such as symbiont mediated defense (found in sweet potato whiteflies infected with P. syringae) or fecundity compensation (found in pea aphids infected with P. syringae). A current project in the lab is focused on aphid avoidance of virulent bacteria. We have found that pea aphids can avoid some virulent strains in the phyllosphere, and therefore avoid infection. The goal of this project is to understand how aphids identify virulent bacteria and which bacterial traits are important.
Breadth of effects
In work funded by the USDA, we aim to understand how diverse aphid species are influenced by P. syringae, specifically how many aphid species can become infected by the bacteria, and how variables such as bacteria strain, aphid species, and plant host influence these interactions. We are focusing on aphid species that are local crop pests in hopes of understanding how these interactions may be used to control pests.
Complex interactions between bacteria, plants, and the ecological community
 Aphids, parasatoid wasps and mutualist ants on a leaf.
Aphids, parasatoid wasps and mutualist ants on a leaf.
A leaf surface can exhibit a dynamic community of interacting bacteria and insects, including herbivores, predators, parasites and mutualists. We use laboratory and mesocosm experiments to determine how these interactions have direct and indirect effects on different species. For instance, we have found that lady beetle larvae develop more slowly when fed on aphids infected with P. syringae, and that the insect populations on plants can influence bacterial abundance and distribution. We have also begun to use field collections to study these interactions in nature.
Genomic evolution of bacteria in bioluminescent symbiosis
 Flashlight fish, Anomalops katoptron
Flashlight fish, Anomalops katoptron
In her dissertation work Tory focused on genomic evolution of the unculturable luminous bacterial symbionts of flashlight fish. Compared to other bioluminescent bacteria, such as the squid symbiont A. fischeri, flashlight fish symbionts have reduced genomes with limited metabolic capabilities and are likely to be obligately dependent on their hosts. Since flashlight fish symbionts are nested within clades of bacteria that are free-living, it is likely that the evolution of obligate dependence in these bacteria is the result of the host interaction, possibly caused by the behavior or specific environment of the fish host. Although these symbionts have reduced genomes that are very similar to the obligate endosymbionts of insects, they have retained many genes associated with chemotaxis and motility, which are almost universally lost in obligate symbionts, and may be used by anomalopid symbionts for transmission.

Genome reduction has evolved convergently in many different lineages of host-dependent bacteria. However, typically these bacteria are physically restricted (intracellular) to hosts and vertically transmitted through host generations. Unusual symbionts such as the flashlight fish symbionts raise of the question of how and why genome reduction might occur when bacteria are not restricted to hosts. To further address this question, we are currently studying a distinct lineage of bacteria that engage in bioluminescent symbiosis with deep sea anglerfish. This project is in collaboration with researchers through the DEEPEND Consortium. Through genomic comparisons with flashlight fish symbionts and well studied symbiont systems we aim to understand the process of genome reduction in extracellular bacteria.
Collaborations
Bacteria are everywhere and Dr. Hendry is broadly interested in all of them. This means the lab is open to collaborations in different systems. We are currently collaborating with other labs on microbiome analyses and are always interested in developing these relationships.
Bacteria are everywhere and Dr. Hendry is broadly interested in all of them. This means the lab is open to collaborations in different systems. We are currently collaborating with other labs on microbiome analyses and are always interested in developing these relationships.
